Recently, I broached the subject of small RNAs, and mentioned that there were two general classes of these regulatory RNA – microRNAs (miRNAs) and short interfering RNAs (siRNAs). These two classes of small RNAs recognize their targets by standard (Watson-Crick) base pairing. However, there are interesting differences regarding target recognition. siRNAs form duplexes that extend the length of the siRNA; this is not unexpected, as the siRNA is derived from a perfectly base-paired precursor that is usually derived from the target itself. miRNAs are a rather different beast; this is the subject of this essay.
First, a brief summary of the means by which miRNAs are produced. In contrast to siRNAs, miRNAs are processed from single-stranded precursors that possess stem-loops that are punctuated with characteristic unpaired bases, or bulges (see Fig. 1). This precursor is processed into small double-stranded RNAs that are largely base-paired, but with a few mismatches. One of these two strands is selectively incorporated into the so-called RNA-induced silencing complex (or RISC), the “business end” of the process. This strand in turn base pairs with its target, leading to any of the outcomes listed in the earlier essay. It is this last step, the actual pairing of miRNA and target, that adds so much flexibility and possibility to regulation by miRNAs.
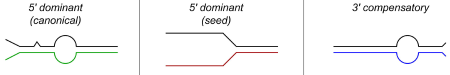
While miRNAs are 22 nts in length, it turns out that the hybrids they form with their targets may be much shorter. Many specific cases have been studied, and numerous artificial examples analyzed, and a relatively straightforward picture has emerged. This is shown in Fig. 2 (which has been lifted from the paper by Brennecke et al., 2005). Basically, a minimal sequence (or “seed”) of 7 nts at the 5’ end of the miRNA suffices for targeting and subsequent effects; this has been shown by introducing such minimal targets into test RNAs (among other ways). This seed can be supplemented by additional pairing at the 3’ end. Alternatively, a small 5’ homology can be “strengthened” by more robust pairing at the 3’ end.
The remarkable part of this is the relatively small size of the seed – 7 nts. Any given 7-mer will occur at random once every 16,000 bases or so; this translates into about once every 10 or so different mRNAs (roughly speaking, with liberal rounding off to make the typing easy). Obviously, a given miRNA does not control 10% of a typical transcriptome (I apologize for this word, but I cannot come up with a concise alternative). But genome-scale studies have shown that a given miRNA may control the expression of many (hundreds of) mRNAs, and have revealed excellent correlations of miRNA-mediated regulation (revealed by miRNA-dependent changes in expression) with the presence of the corresponding 7 nt target sequence.
This feature has fascinating implications for genome and regulatory evolution. Some of these (there are many, and the subject can fill up a lengthy review article):
- It turns out that the miRNA complement of a cell or organism plays a significant role in sculpting the sequences of mRNAs – abundant and regulated mRNAs (as groups) are disposed to lack targets for miRNAs that are expressed at the same time, while mRNAs of low abundance (such as those that are diminished in a developmental manner) are more likely to possess targets of the complement of miRNAs present in the corresponding cell or tissue.
- Often, miRNA targets are conserved amongst related organisms. In such cases, as a rule, the target sequences seem to be under so-called purifying selection; in other words, they are less variable than surrounding sequences when compared with their counterparts in closely-related organisms. This suggests that natural selection acts to preserve this mode of regulation.
- Conversely, many miRNA targets are not conserved, but rather are species-specific. This evolutionary flexibility probably reflects the small size of the target sequence seed; in other words, new targets may arise with a relatively high frequency, and existing targets (especially those not fixed by selection or the further creation of 3’ anchors) may occasionally be lost due to random mutation.
- It might seem, at first glance, that the prospect of an underworld of small regulatory RNAs might be a way by which one might reduce or eliminate the “junk DNA problem” (which is, in a nutshell, the prospect that eukaryotic genomes are rife with so-called junk). It seems reasonable to suppose that small RNAs, that obviously may control many eukaryotic genes, could make a large dent in the quantity of junk DNA seen in a typical eukaryotic cell. However, the above numbers dispel this notion – a few hundred miRNAs suffice to control (in principle) all genes in a eukaryotic cell. Said another way, a few hundred miRNA genes would constitute no more than a million bp of DNA, an infinitesimally small fraction of the junk DNA in a cell.
Further reading (an admittedly limited sampling):
Brennecke J, Stark A, Russell RB, Cohen SM. 2005. Principles of MicroRNA-Target Recognition. PLoS Biology 3, e85 doi:10.1371/journal.pbio.0030085
Farh KK et al. 2005. The Widespread Impact of Mammalian MicroRNAs on mRNA Repression and Evolution. Science 310, 1817-1821.
Grimson A, Farh KK, Johnston WK,Garrett-Engele P, Lim LP, Bartel DP. 2007. MicroRNA Targeting Specificity in Mammals: Determinants beyond Seed Pairing. Mol. Cell 27, 91-105.

You are so awesome! I don’t think I’ve read through anything like this before.
So wonderful to discover another person with genuine
thoughts on this subject. Seriously.. thank you for
starting this up. This website is something that’s needed on the web, someone
with some originality!
link: anything new? absolutely ove this thank you so much